Database of curated and re-analyzed gene expression studies
Gemma provides data, experimental design annotations, and differential expression analysis results for
thousands
of microarray and RNA-seq experiments. We re-analyze raw data from public sources (primarily NCBI GEO),
annotate experimental conditions, conduct quality control and compute differential expression using
standardized
procedures. We have especially good coverage of experiments relevant to the nervous system. See the documentation for more information.
Gemma was and developed and is maintained by the Pavlidis
group at UBC.
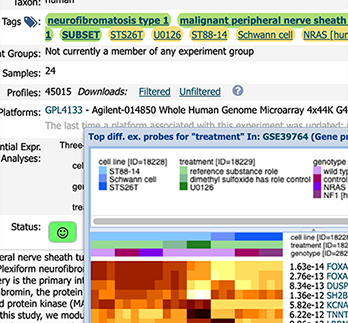
Convenient programmatic access to Gemma's data and analyses is available via the software packages gemma.R (R/Bioconductor) and gemmapy (Python).
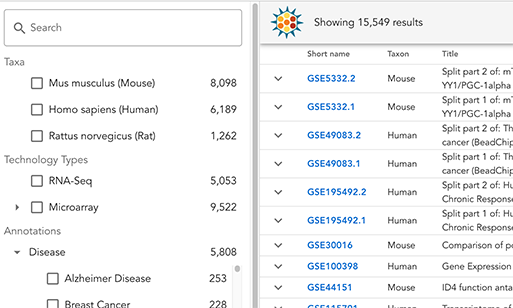
We invite you to try out the new Gemma Browser, our new interface for exploring and searching Gemma's data holdings. It's still in beta, and more features and improvements are planned, but we'd love to hear your feedback.
Questions? Feel free to reach out.

